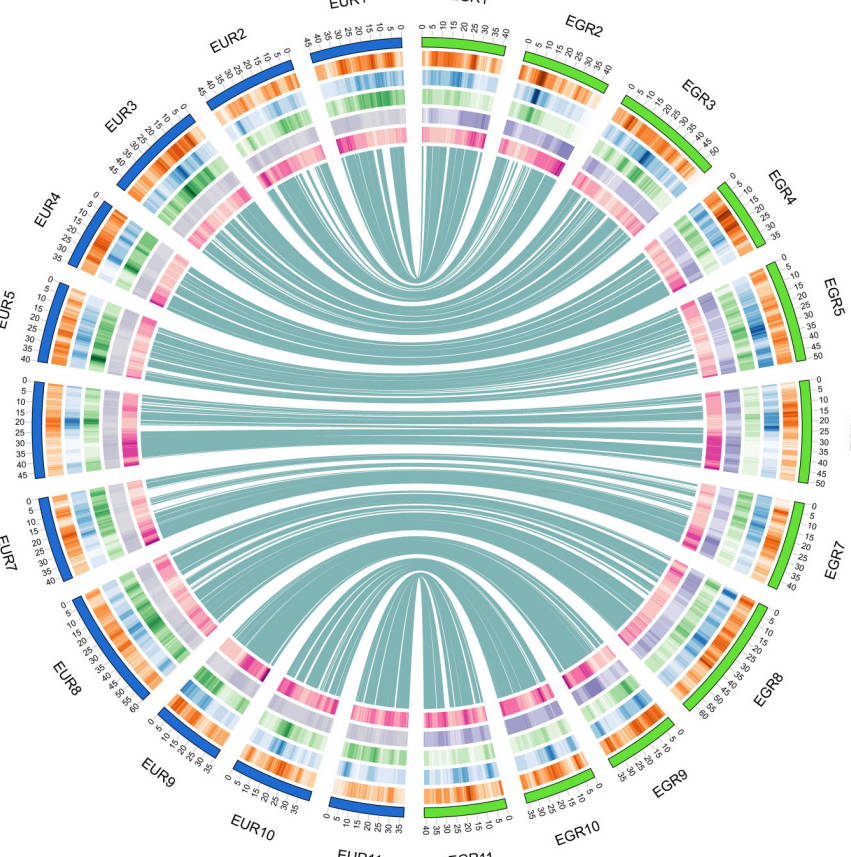
This study constructed a chromosome-level genome assembly of Eucalyptus urophylla × Eucalyptus grandis (E. urograndis) clone DH32-29 using BGI-SEQ500, PacBio-SMRT, and Hi-C technologies, with a genome size of 592.09 Mb and a scaffold N50 of 58.62 Mb.
https://doi.org/10.1186/s12870-025-07371-3Using sequencing technologies such as PacBio HiFi long reads and Omni-C, the researchers successfully constructed a haplotype-resolved reference genome (v4.0) for Eucalyptus grandis TAG0014, showing significant improvements in contiguity and completeness.
https://doi.org/10.1093/g3journal/jkaf112Researchers generated a high-quality chromosome-level genome assembly of Eucalyptus globulus Labill. with a genome size of 556.98 Mb, a contig N50 of 37.93 Mb, and a scaffold N50 of 54.03 Mb, providing a valuable resource for elucidating molecular mechanisms.
https://doi.org/10.1038/s41597-025-05421-xQuick Access
Eucalyptus Genome Research
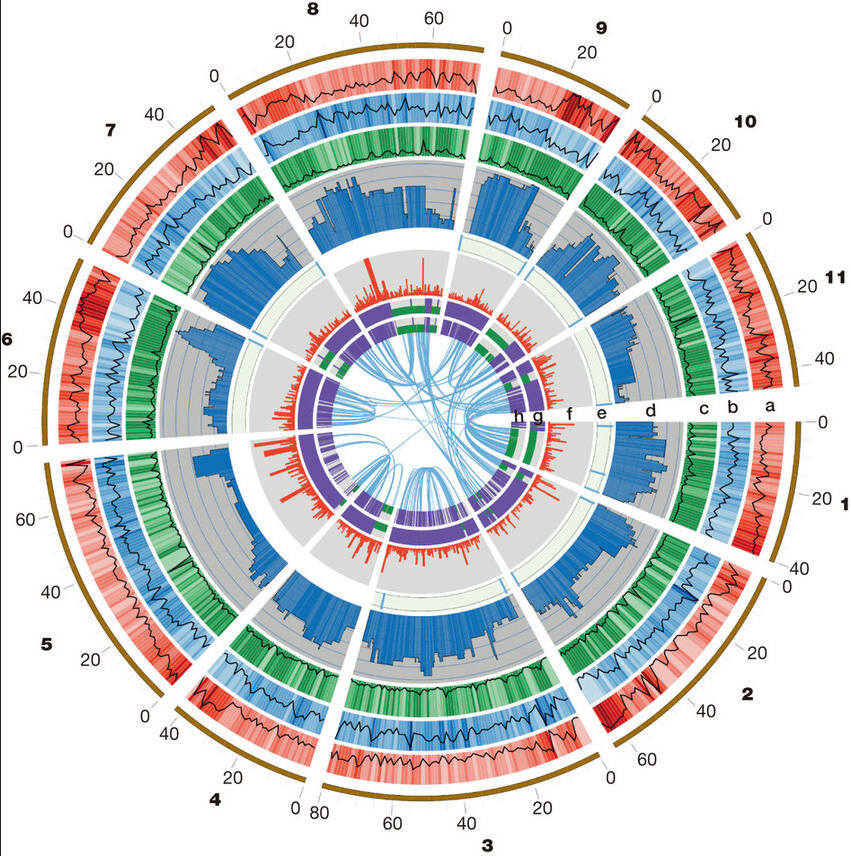
Eucalyptus grandis genome
Link to paper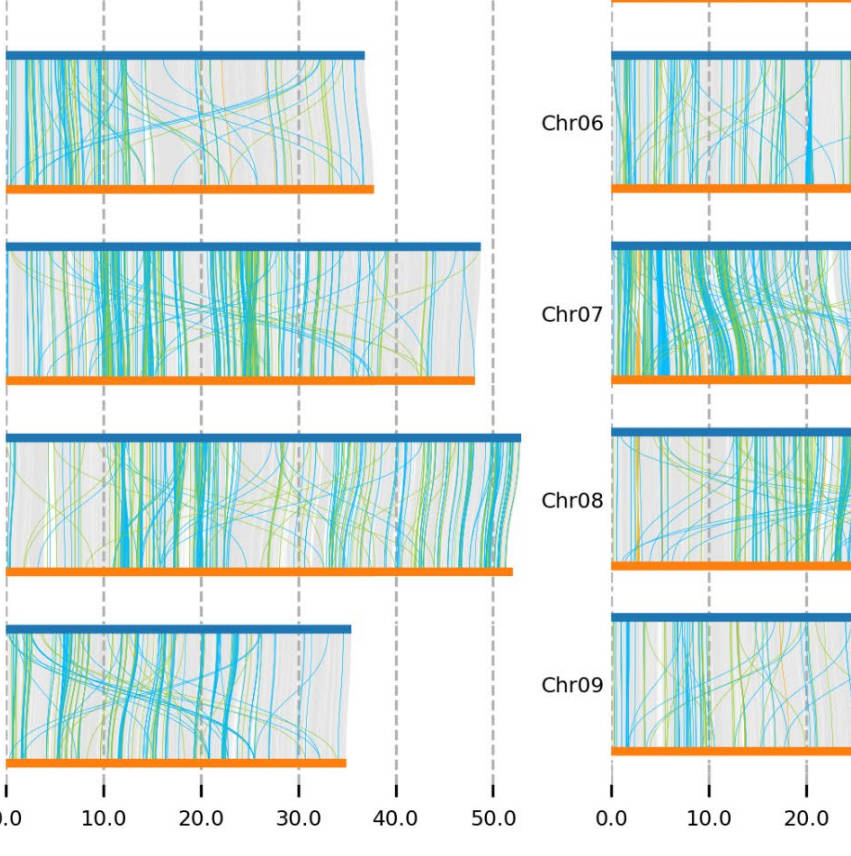
T2T Eucalyptus Genome
Link to paper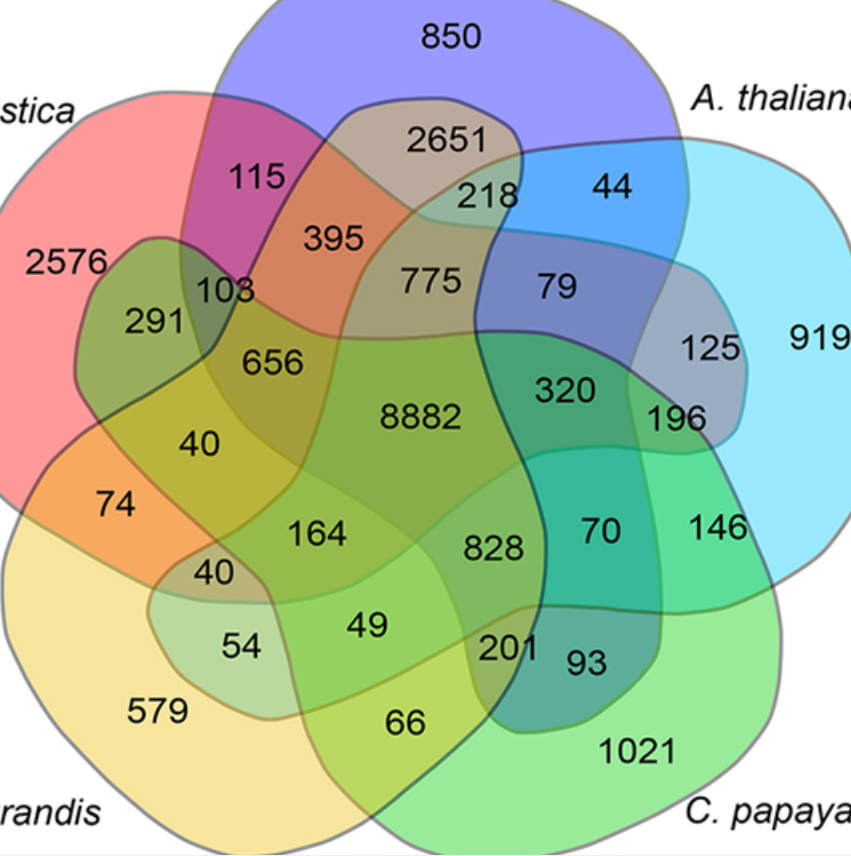
Comparative genomics
Link to paper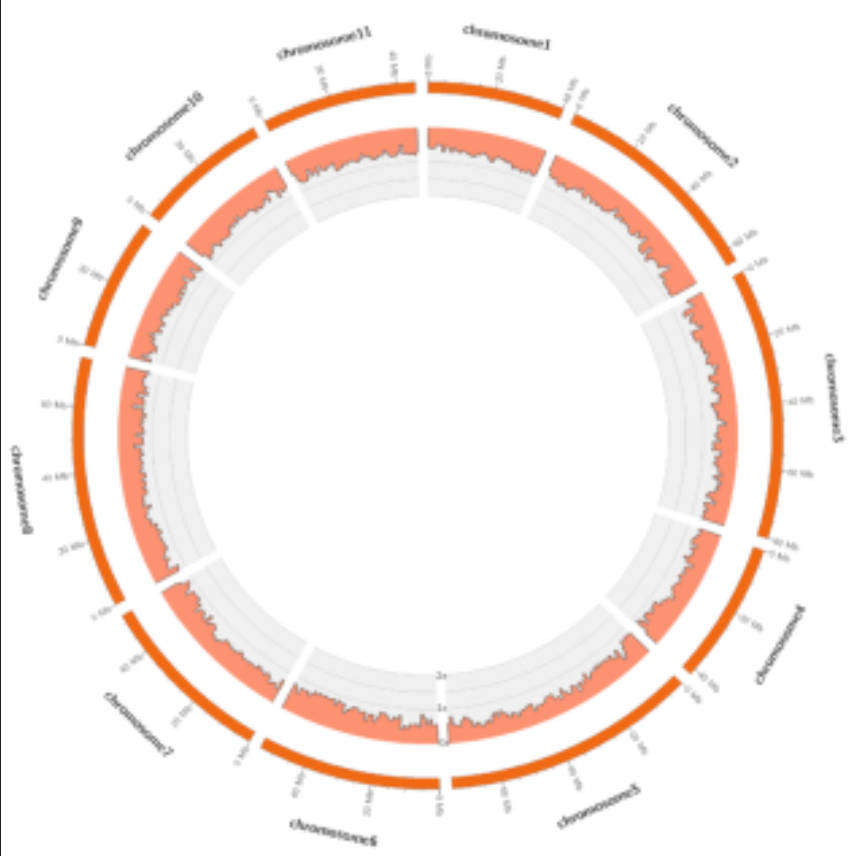
Eucalyptus pauciflora Genome Assembly
Link to paper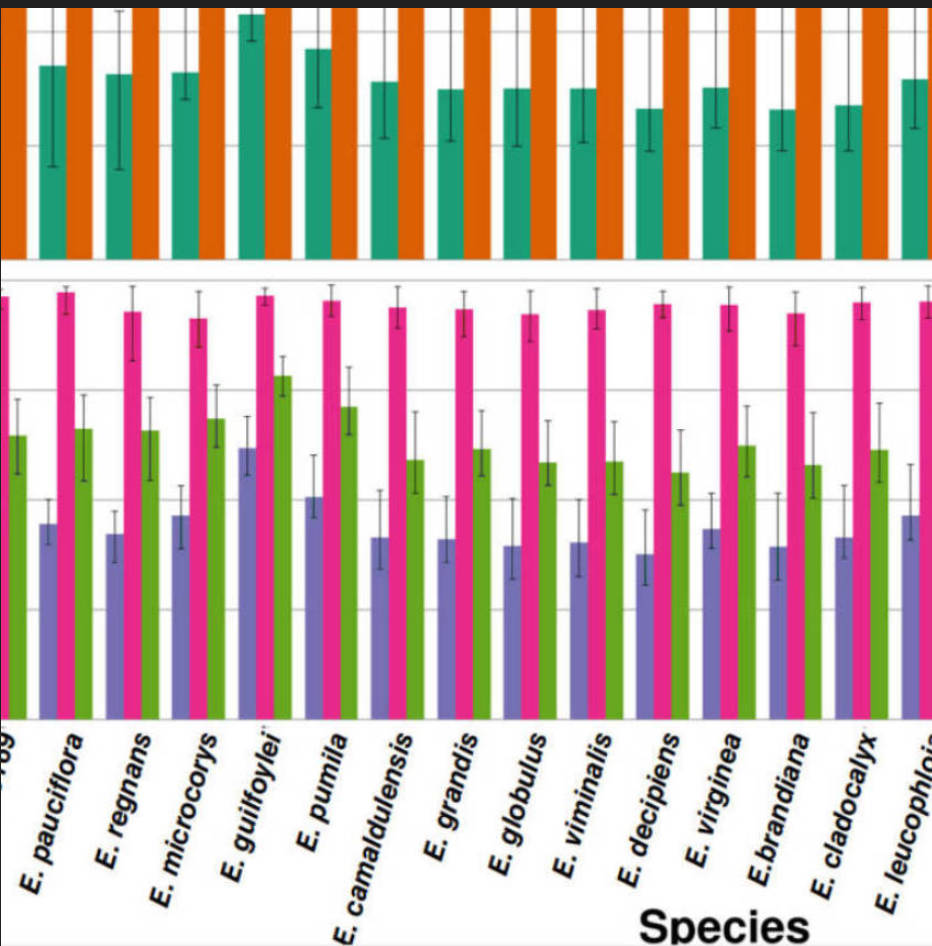
Eucalyptus Genome Evolution
Link to paper