7 Protein Interaction
This tool focuses on protein-protein interaction analysis of Eucalyptus. It integrates multi-source evidence (experiments, literature, genomic predictions, etc.) through the STRING database to construct an interaction network between Eucalyptus proteins. This function can help decipher the molecular mechanisms of core physiological processes such as Eucalyptus growth and stress resistance, providing key support for functional gene research and targeted breeding of Eucalyptus.
7.1 Operation Demonstration
7.1.1 Enter Eucalyptus-Specific Protein IDs
In the "Query Proteins" input box, enter Eucalyptus-specific protein IDs separated by spaces or commas. Example query:
EUGRSUZ_C02720 EUGRSUZ_C02894 EUGRSUZ_D02251 EUGRSUZ_F01229 EUGRSUZ_F03074 EUGRSUZ_H02295 EUGRSUZ_H04045 EUGRSUZ_I00055 EUGRSUZ_I00628
Figure 7.1: Screenshot of Eucalyptus-specific protein ID input interface for Protein Interaction
7.1.2 Set Analysis Parameters
Minimum Confidence Score: This parameter is used to filter the reliability of interaction relationships, providing multiple threshold levels:
- Highest (0.900): Retains only extremely high-quality interactions, resulting in the most streamlined network;
- High (0.700): High-confidence interactions, balancing reliability and network scale;
- Medium (0.400): Medium confidence (default selection), suitable for most analysis scenarios;
- Low (0.150): Low confidence, resulting in the most abundant network;
- Custom Value: Custom threshold to meet special analysis needs.
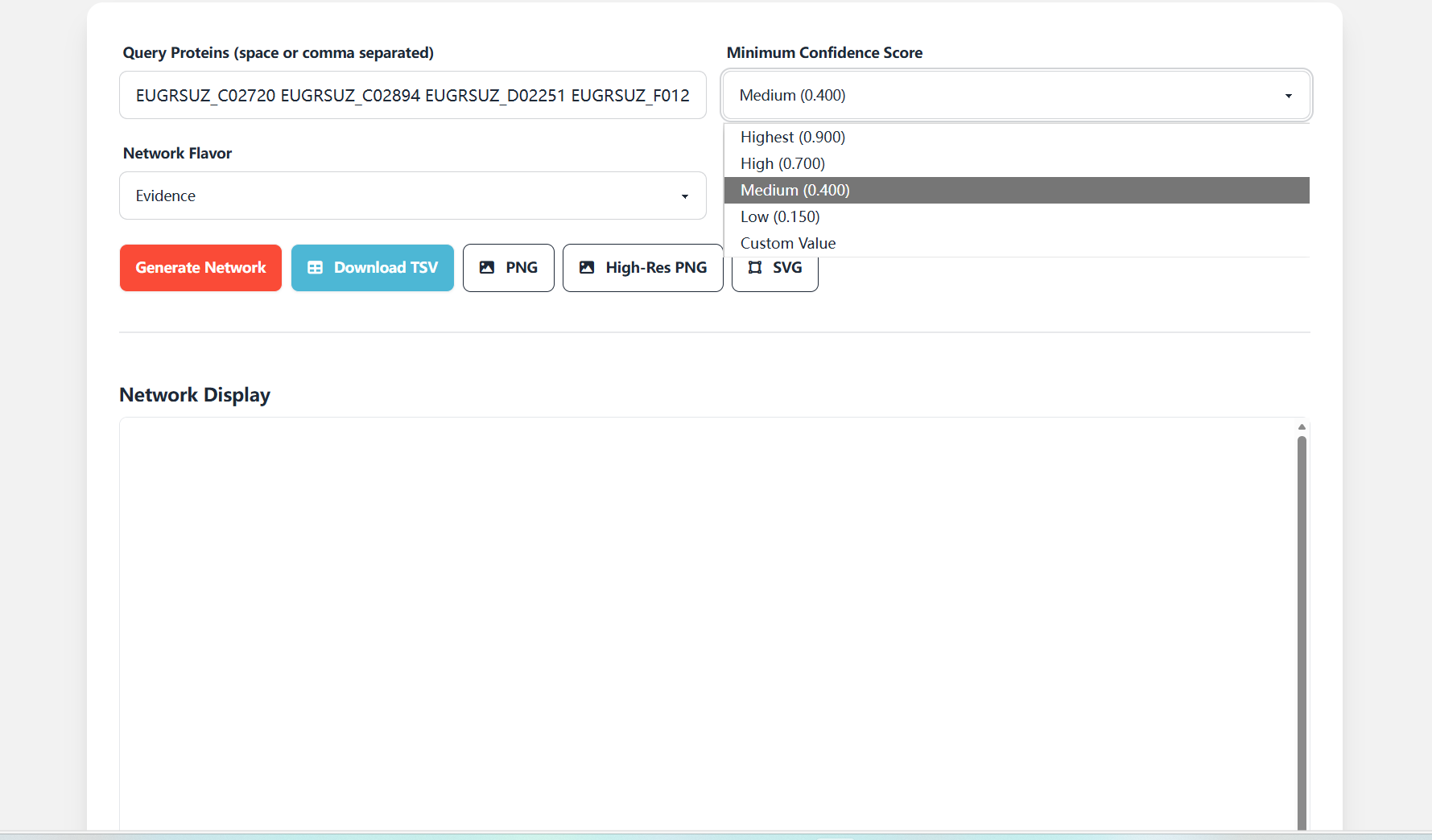
Figure 7.2: Screenshot of Minimum Confidence Score setting interface for Protein Interaction
Figure 7.3: Screenshot of custom Minimum Confidence Score setting interface for Protein Interaction
Network Flavor: Two modes are available for constructing interaction networks:
- Evidence: Integrates multi-source interaction evidence such as experiments, literature, and calculations, suitable for global functional analysis of Eucalyptus proteins;
- Confidence: Constructs the network based on confidence weights, emphasizing the quantitative reliability of Eucalyptus protein interactions.
Figure 7.4: Screenshot of Network Flavor setting interface for Protein Interaction
7.1.3 Generate and Download the Interaction Network
Click the "Generate Network" button to generate a customized Eucalyptus protein interaction network. The platform provides multiple download options for result preservation and academic use:
- Download TSV: Downloads the interaction relationship table, including Eucalyptus protein pairs, confidence scores, and species-specific protein annotations;
- PNG/High-Res PNG: Downloads network visualization images (high-resolution PNG format is suitable for academic presentations and publications);
- SVG: Downloads vector graphics format, supporting unlimited scaling without quality loss.
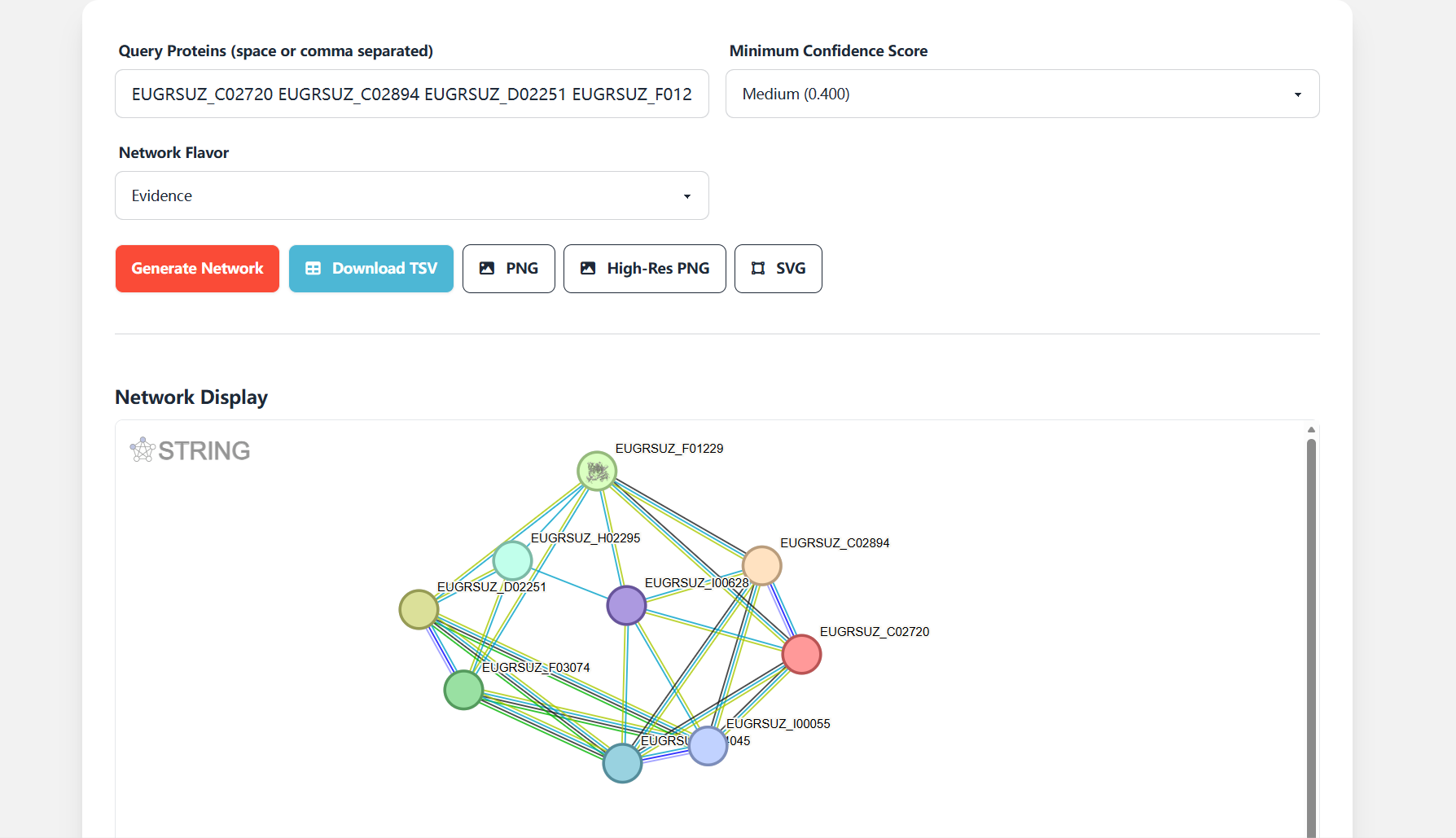
Figure 7.5: Screenshot of interaction network generation and download interface for Protein Interaction
7.2 Result Analysis
Network Visualization Interpretation
The interaction network uses nodes and edges to represent Eucalyptus protein interactions, with clear visual encoding rules:
View Protein Details by Clicking Nodes.After generating the interaction network, click any node to instantly display a pop-up window with exclusive information about the Eucalyptus protein, including three core information modules:
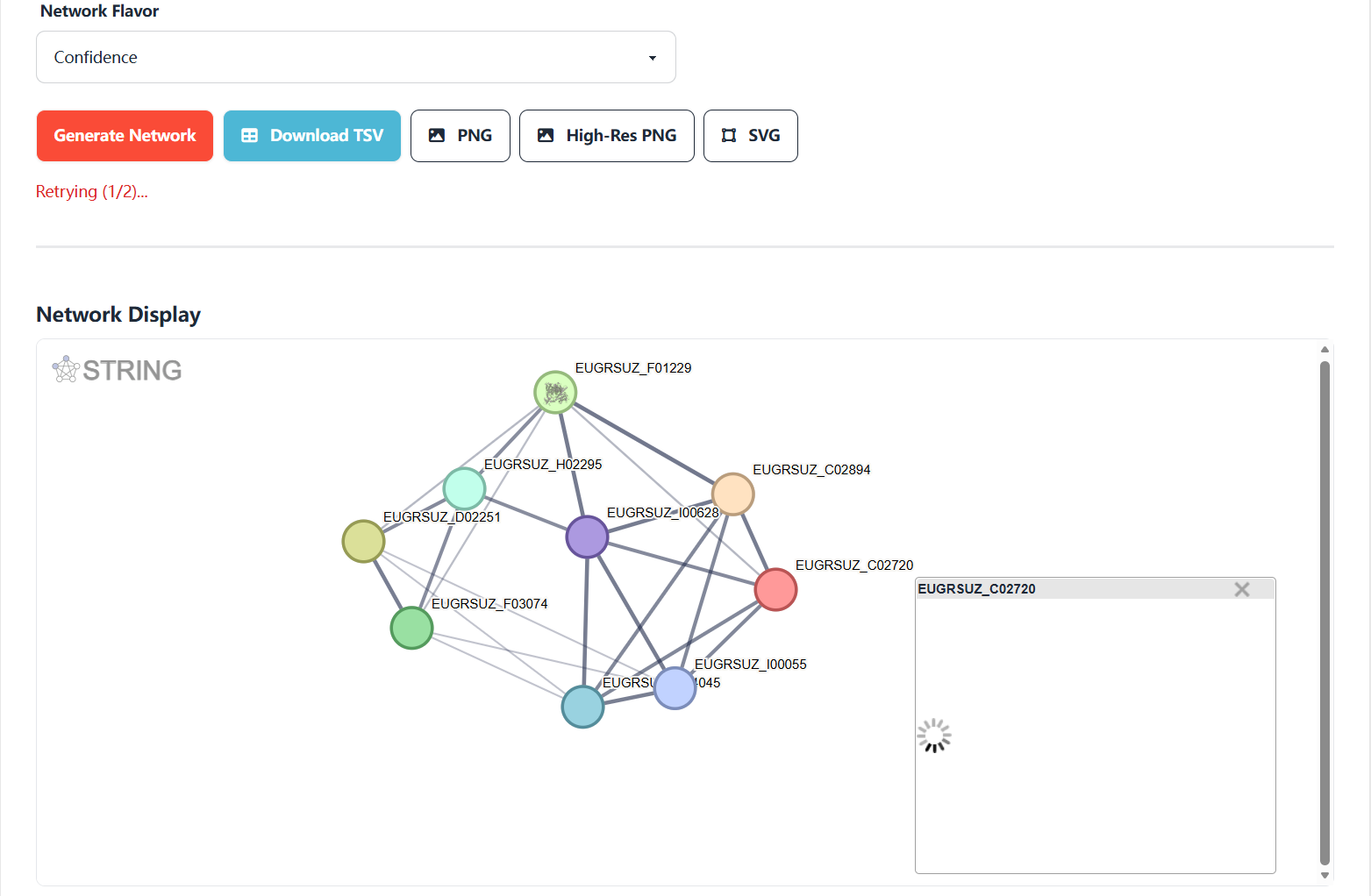
Figure 7.6: Screenshot of Eucalyptus protein interaction network result and detail view
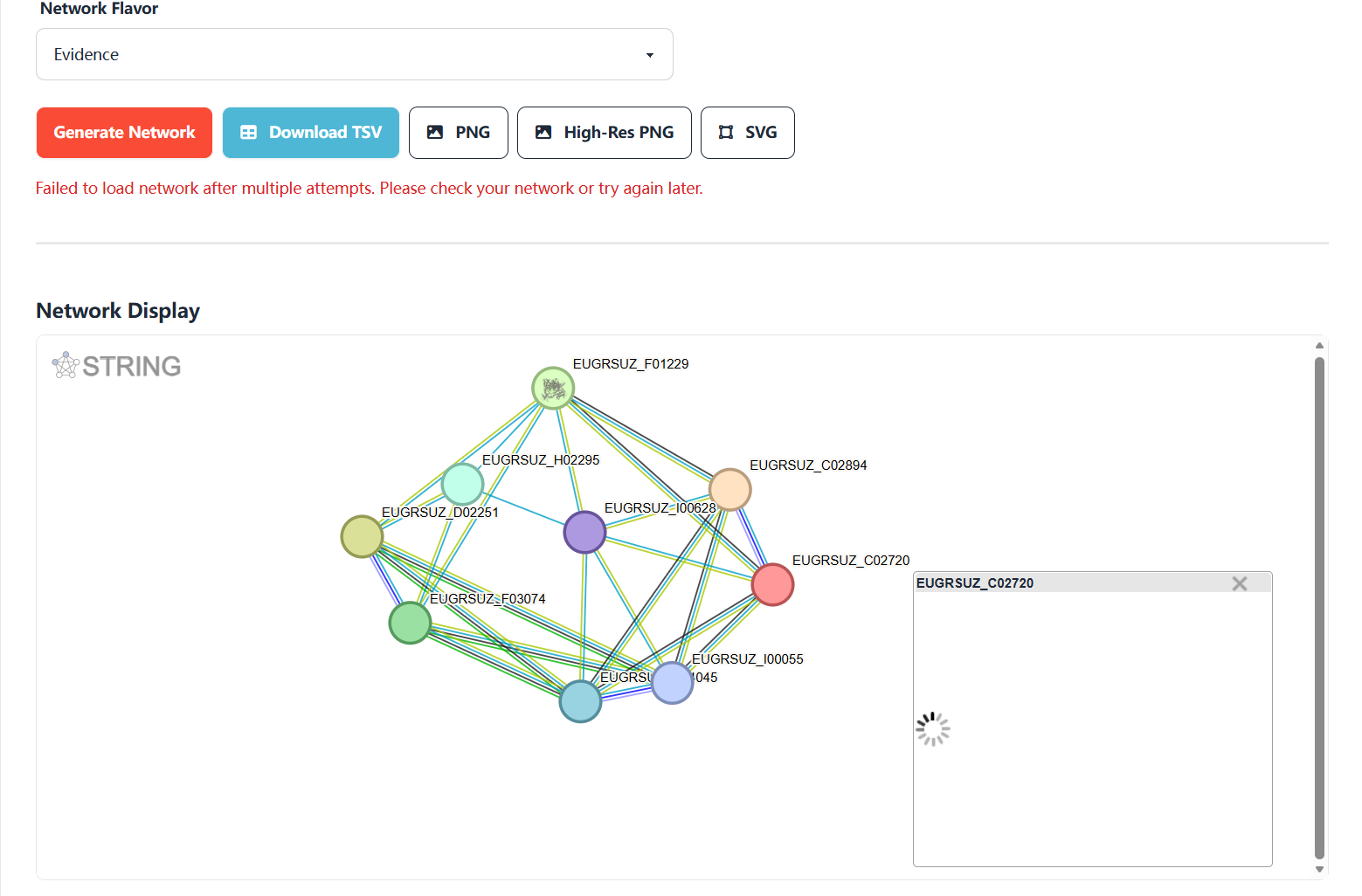
Figure 7.7: Screenshot of Eucalyptus protein interaction network node detail pop-up
Link to STRING Official Website for In-Depth Data
A dedicated "STRING" label is displayed on the network interface (located on the side of the network). Click this label to directly jump to the corresponding page of the STRING official database to obtain more comprehensive in-depth data for Eucalyptus protein research:
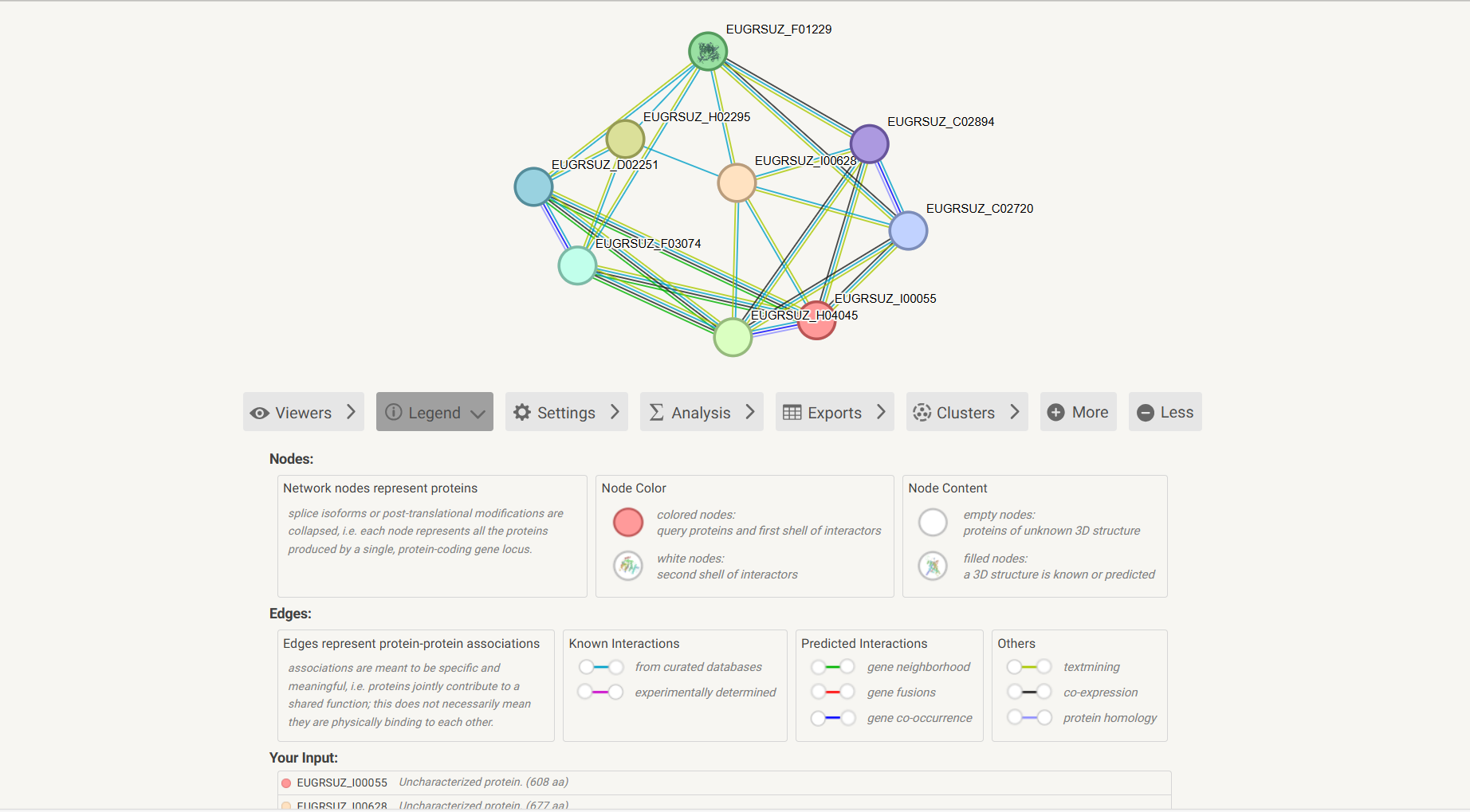
Figure 7.8: Screenshot of STRING official website link entry for Protein Interaction