4 VCF Browse
VCF Browser is a core retrieval and research support tool for data from the eucalyptus variation detection project, focusing on the precise detection and in-depth research needs of variant sites. It centrally collects extensive data from the eucalyptus variation detection project, enabling standardized storage and management of data, and provides accessible variant site resources for researchers. By integrating multi-dimensional filtering criteria such as chromosomes, position ranges, and VCFIDs, it helps users quickly locate variant sites of interest. Meanwhile, it supports exporting query results in CSV format, which is compatible with subsequent further research on these sites (e.g., functional verification, mechanism analysis, association studies, etc.). With efficient site retrieval capabilities, this tool provides accurate data support for researchers throughout the entire process from variant site screening to in-depth research, serving as a core data hub for conducting variation-related molecular biology research.
4.1 Operation demonstration
- Mode One Genome-Wide Chromosome Data Retrieval: Obtain all variant data of a complete chromosome in the target data table.
- Mode Tow Genomic Position Interval Retrieval: Obtain variant data within a specific base position range on a chromosome.
- Mode Three VCFID Exact Search: Locate the data of a single variant with a known unique identifier (VCFID).
4.1.1 Genome-Wide Chromosome Data Retrieval
Select the data table: Click the Select Table dropdown menu to choose the target data table.
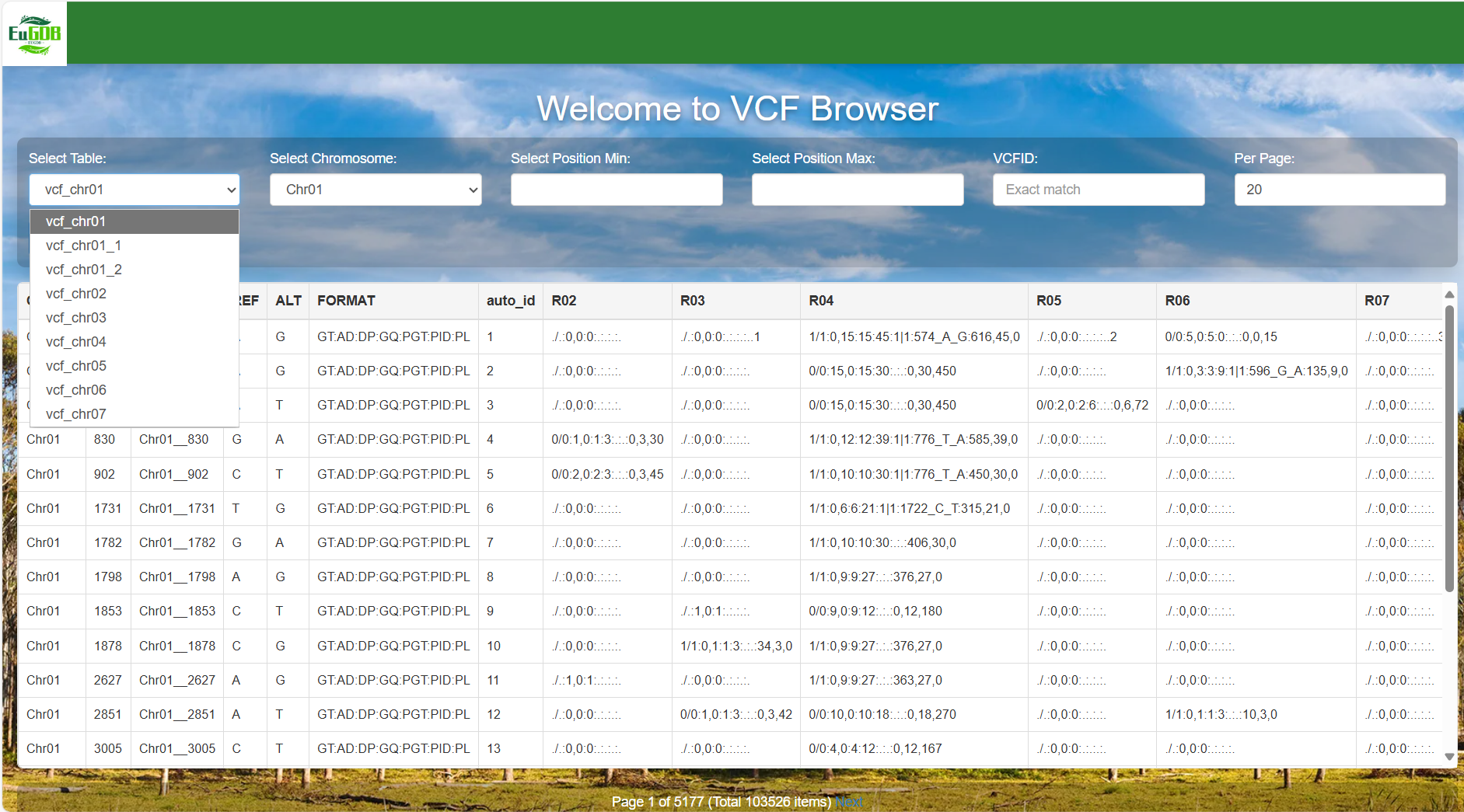
Figure 4.1: A screenshot of the one type of selection for VCF Browser.
Select the chromosome: Click the Select Chromosome dropdown menu to select the target chromosome.
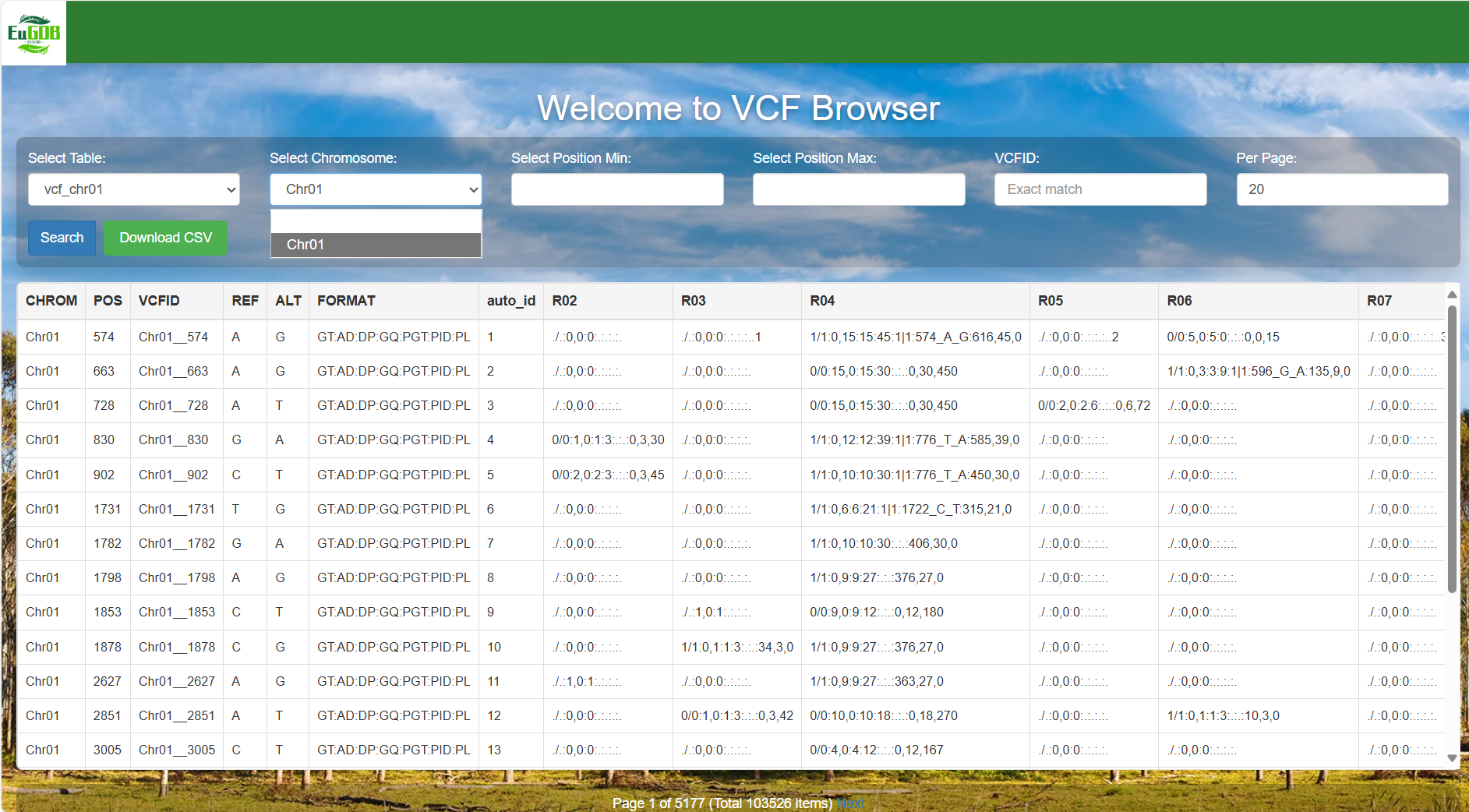
Figure 4.2: A screenshot of the one type of selection for VCF Browser.
Set the number of items per page and execute retrieval: Enter the number of items to be displayed per page in the Per Page input box, then click the Search button to view all variant data of the selected chromosome in the chosen table.
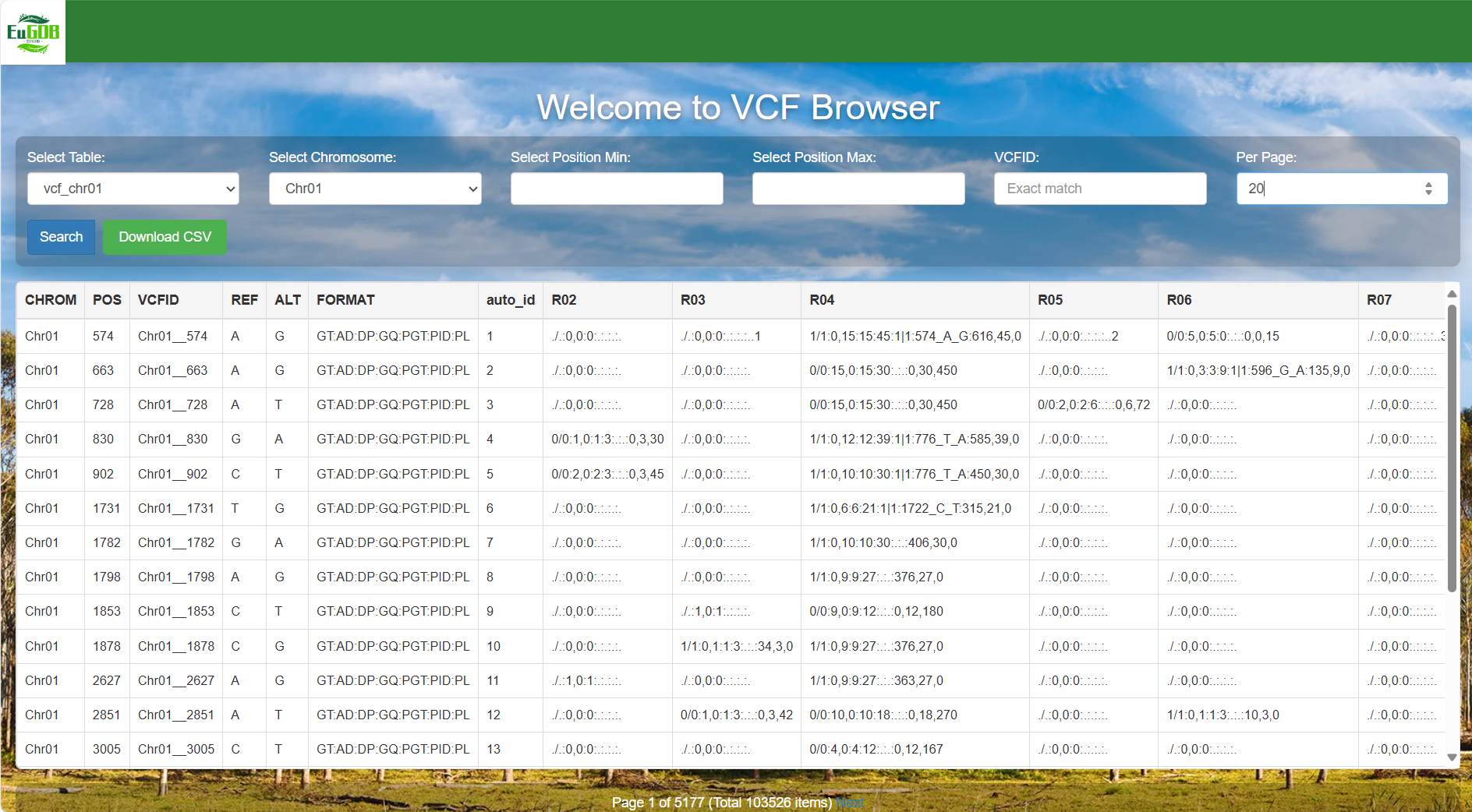
Figure 4.3: A screenshot of the one type of selection for VCF Browser.
4.1.2 Genomic Position Interval Retrieval
Select the data table: Click the Select Table dropdown menu to choose the target data table.

Figure 4.4: A screenshot of the one type of selection for VCF Browser.
Select the chromosome: Click the Select Chromosome dropdown menu to select the target chromosome.

Figure 4.5: A screenshot of the one type of selection for VCF Browser.
Set the position range: Enter the start position (e.g., 1000) in the "Select Position Min" input box. Enter the end position (e.g., 3000) in the "Select Position Max" input box.
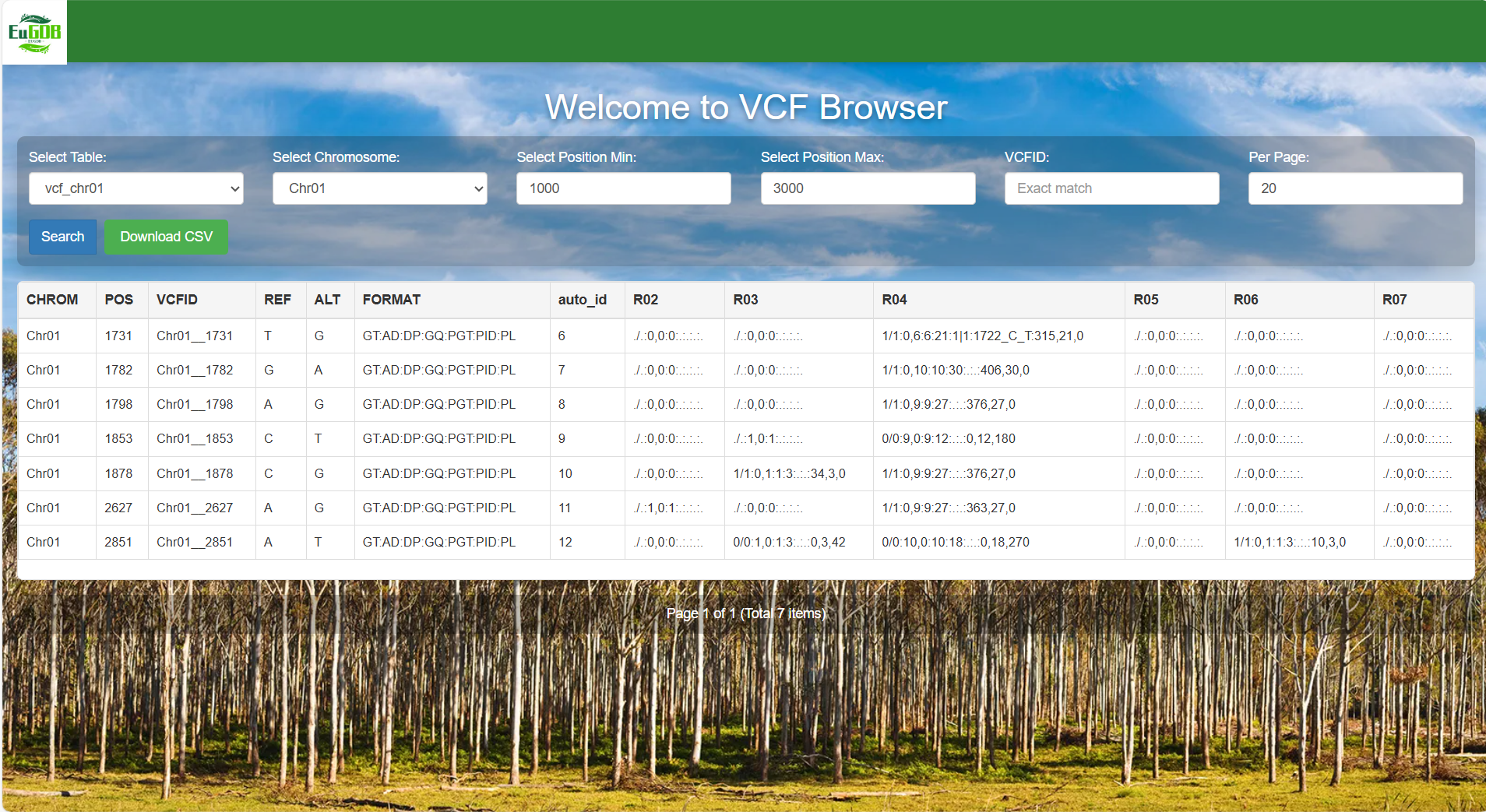
Figure 4.6: A screenshot of the one type of selection for VCF Browser.
Set the number of items per page and execute retrieval: Enter the number of items to be displayed per page in the Per Page input box, then click the Search button to view all variant data of the selected chromosome in the chosen table.
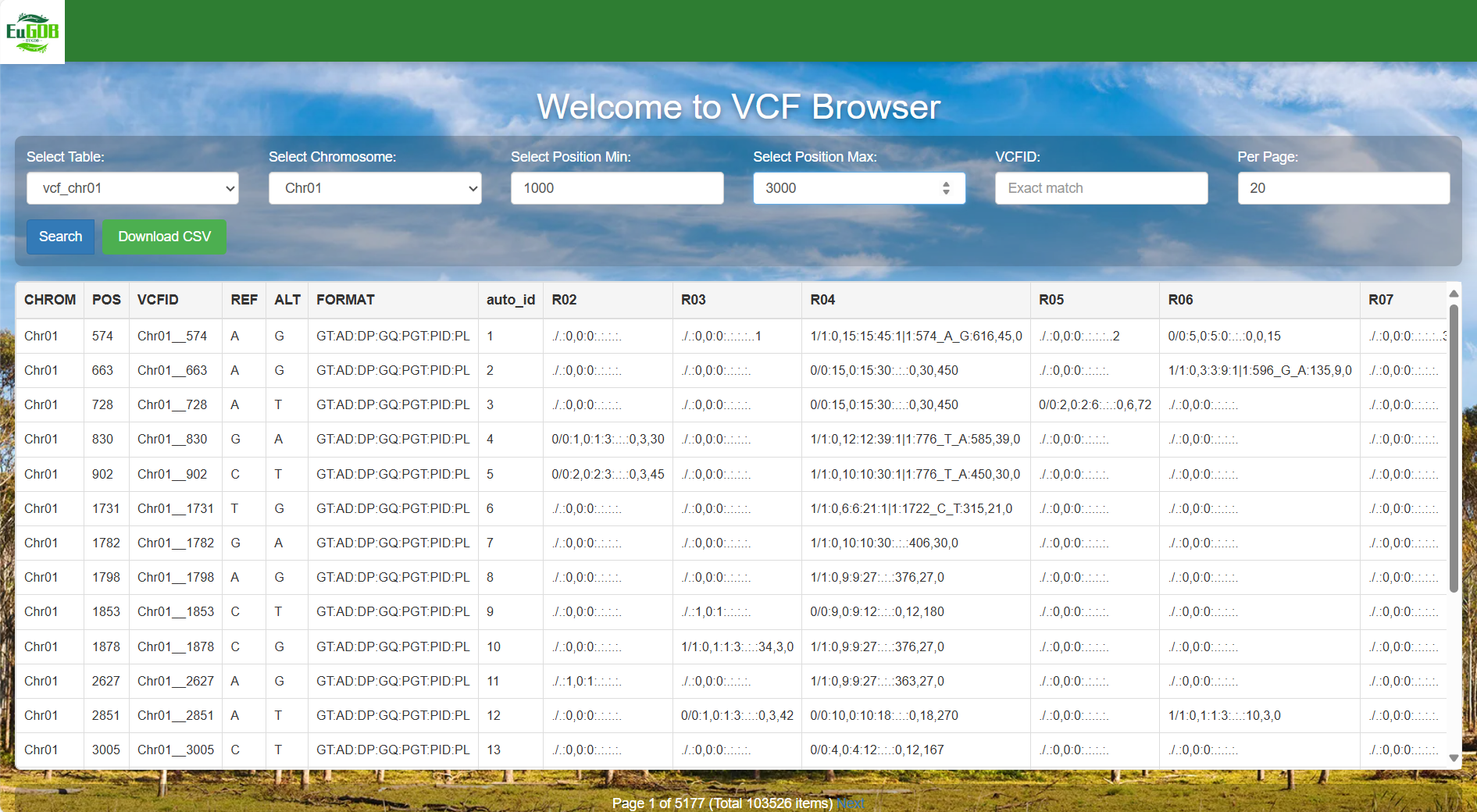
Figure 4.7: A screenshot of the one type of selection for VCF Browser.
4.1.3 VCFID Exact Search
Select the data table: Click the Select Table dropdown menu to choose the target data table.

Figure 4.8: A screenshot of the one type of selection for VCF Browser.
Select the chromosome: Click the Select Chromosome dropdown menu to select the target chromosome.

Figure 4.9: A screenshot of the one type of selection for VCF Browser.
Enter the VCFID: Input the unique ID of the target variant (e.g., Chr01_830) in the "VCFID" input box.
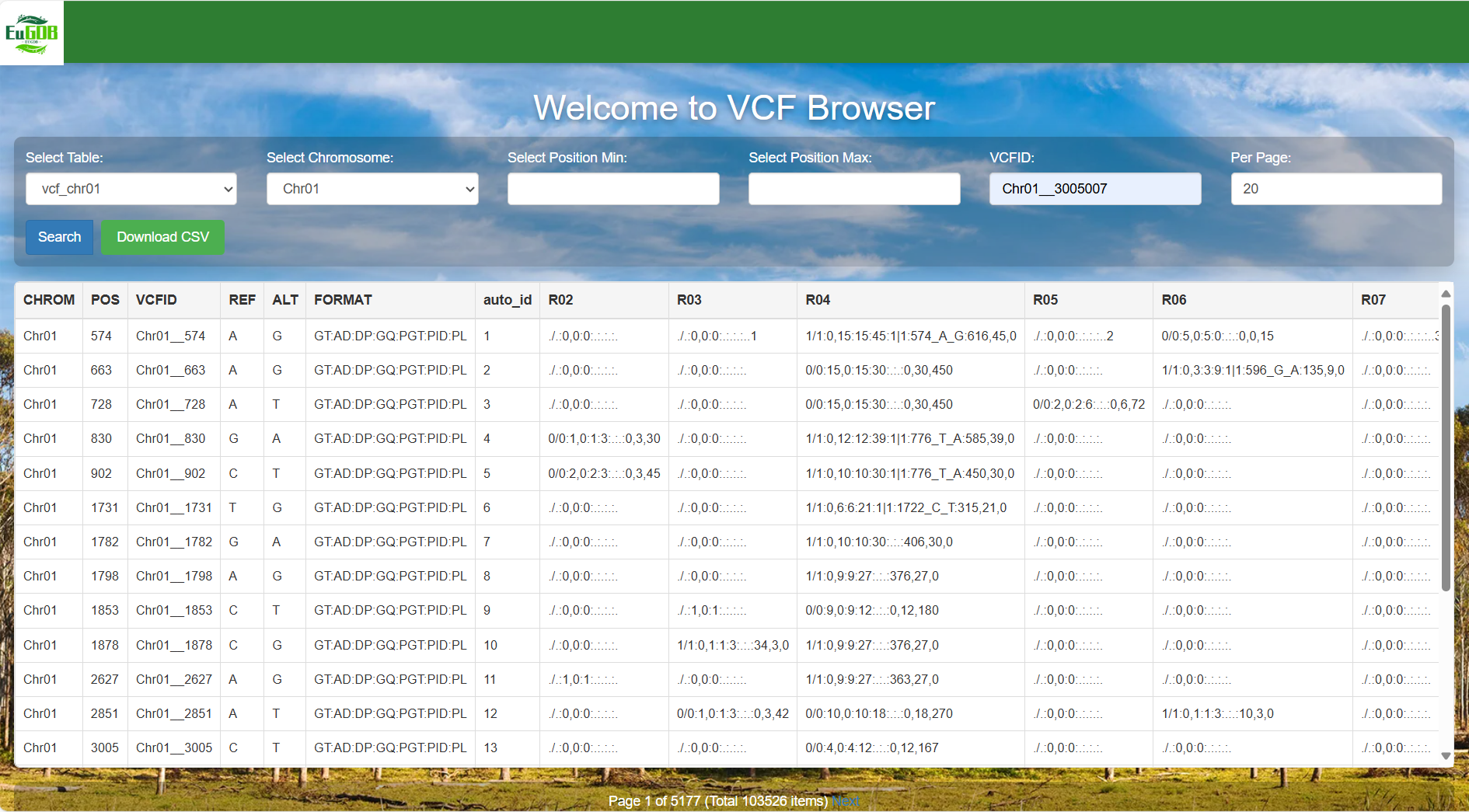
Figure 4.10: A screenshot of the one type of selection for VCF Browser.
Set the number of items per page and execute retrieval: Enter the number of items to be displayed per page in the Per Page input box, then click the Search button to view all variant data of the selected chromosome in the chosen table.
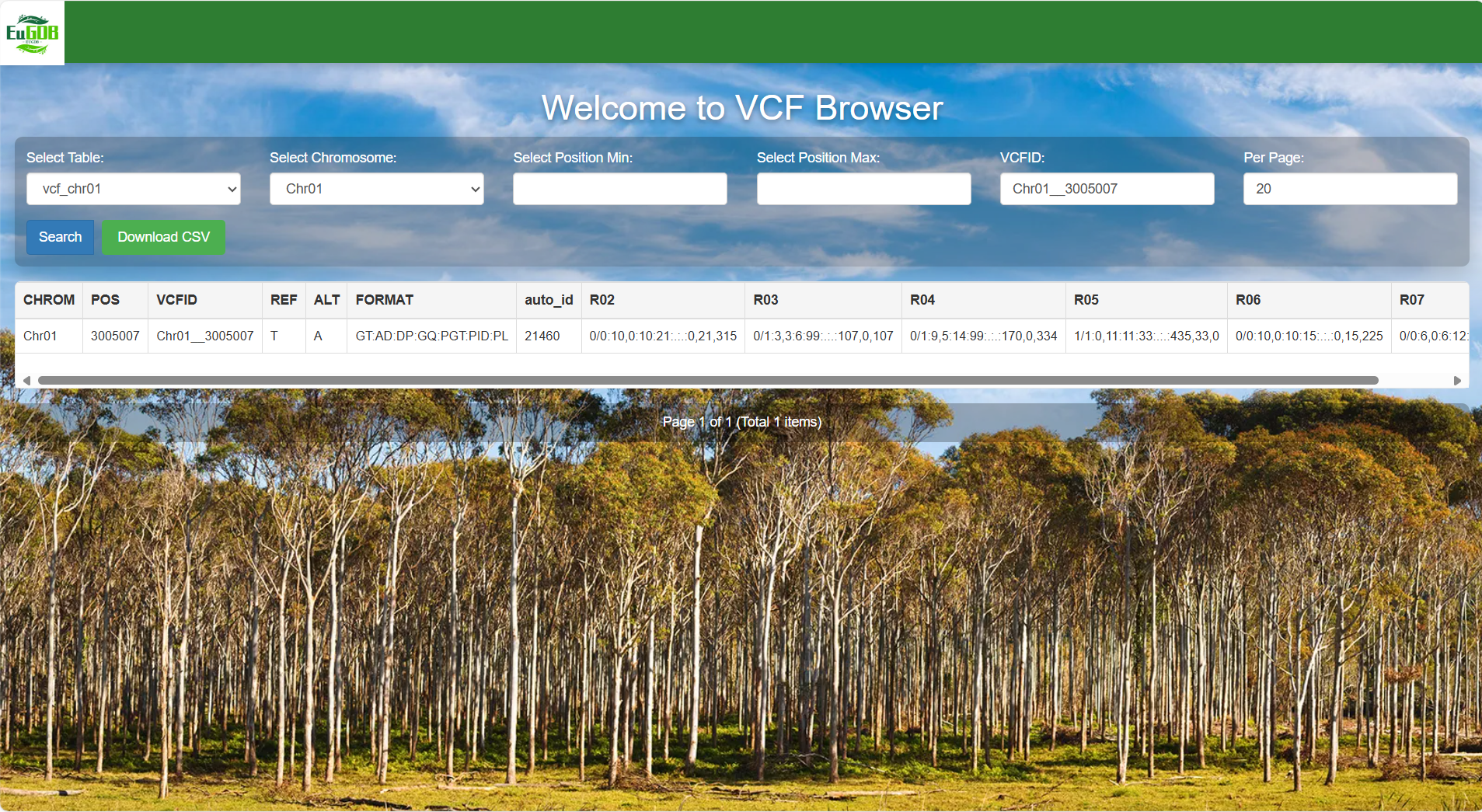
Figure 4.11: A screenshot of the one type of selection for VCF Browser.
4.2 Result interpretation
The chromosome where the variant is located. For example, "Chr01" indicates that the variant is on chromosome 1.
The genomic coordinate position of the variant on the chromosome. For example, "574" indicates that the variant is at the 574th base position on the chromosome.
The unique identifier of the variant, such as "Chr01_574", facilitating subsequent tracing and citation of the variant.
The reference allele of the reference genome, i.e., the base type of the locus in the reference genome (e.g., "A").
The alternative allele after variation, i.e., the base type of the locus after variation (e.g., "G").
Defines the format of variant information in the subsequent sample columns (R02~R07).
Variant information of different samples, whose format corresponds one-to-one with the definition in the FORMAT column.
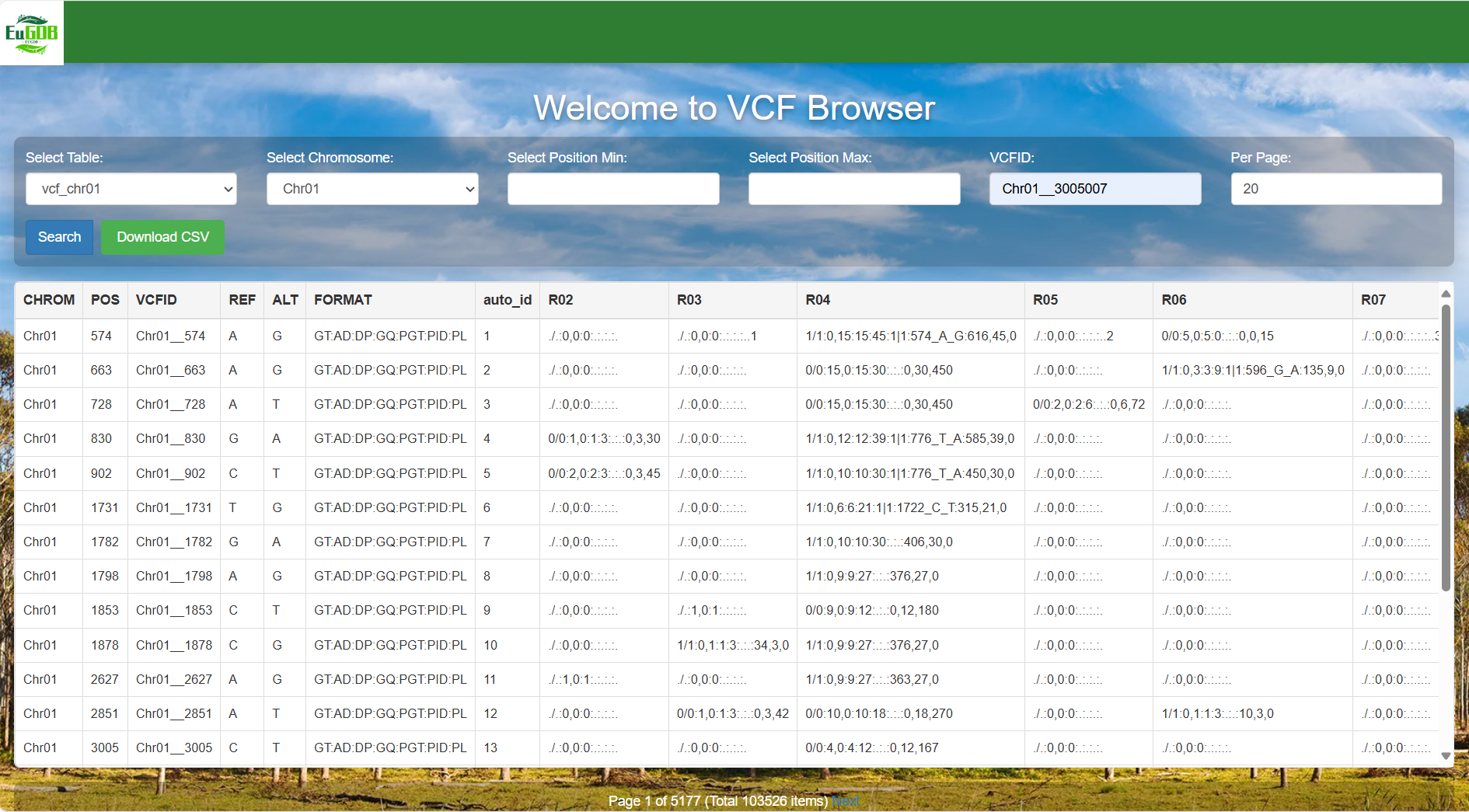
Figure 4.12: A screenshot of the VCF Browser result display.